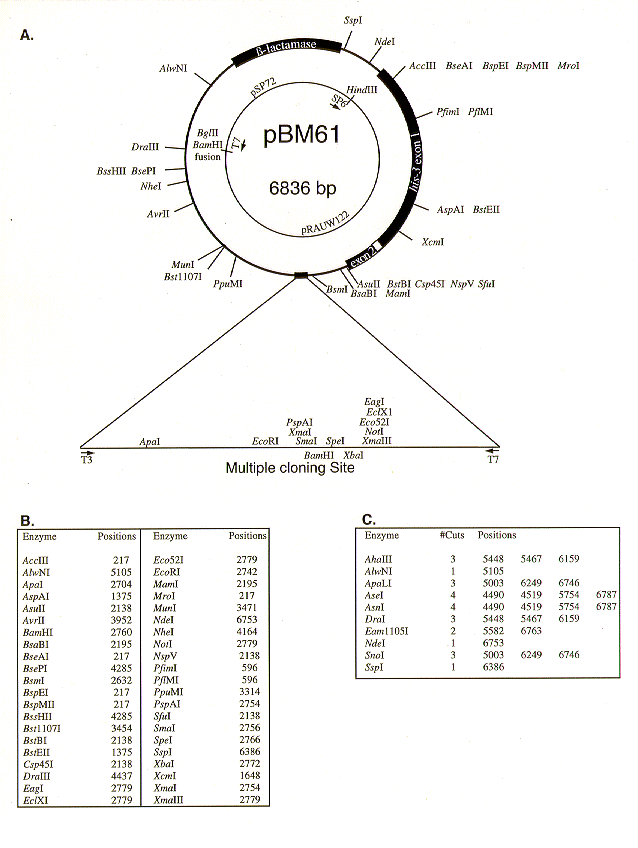
Figure 1. Map and restriction sites of pBM61.
Margolin, B.S., Freitag, M., and Selker, E.U. - Institute of Molecular Biology,
University of Oregon, Eugene, OR 97403 USA
Errata in
FGN 47:112
We report two new plasmids, pBM60 and pBM61, and procedures to efficiently generate single- copy transformants targeted to the his-3 locus in Neurospora crassa.
Gene replacement is a powerful tool to compare properties of different DNA sequences integrated at a common locus in otherwise isogenic strains. In Neurospora crassa, transformation by homologous recombination is normally much less common than transformation by non-homologous recombination. It is therefore desirable to be able to select, positively, for gene-replacement events. Two such methods have been described. The first, developed for the am locus of Neurospora (Miao, V.P., M.J. Singer, M.R. Rountree and E.U. Selker, 1994 Mol. Cell. Biol. 14: 7059-7067) has the feature that it typically yields homokaryotic transformants, but the efficiency of the method is sometimes low (unpublished observations). The second system, developed for the his-3 locus of Neurospora (Aramayo and Metzenberg, 1996, Fungal Genet. Newsl. 43: 9-13) is somewhat more efficient, but still does not give a large number of transformants. We encountered other difficulties with this system that led us to modify it. The his-3 targeting plasmid pRAUW122 is rather large (14.3 kb) and therefore can be difficult to work with. In experiments with pRAUW122 or its derivatives, we found that it was not necessary to screen for benomyl- sensitive strains, as the number of benomyl- resistant transformants obtained without selection on benomyl was low (circa 5%). To render the large plasmid more compact, we therefore deleted the Bml gene and some of the his-3 3' sequences from pRAUW122. In addition, using pRAUW122 derivatives, we frequently found His+ transformants that showed wild-type his-3 bands in Southern analyses but did not have the test sequences targeted to his-3. Because mock transformations never resulted in His+ transformants, we assumed that His+ transformants were products of recombination events occurring solely within the 5.22 kb 5' his-3 region of pRAUW122, thereby restoring his-3 function but excluding the test sequences. We therefore inserted cloning sites closer to the 5' end of the truncated his-3 gene. To increase the number of useful cloning sites, the pBluescript KS(+) multiple cloning site (Stratagene, La Jolla, CA) was used in place of the original four cloning sites (see Figure 1).
Methods. We created two compact targeting plasmids, pBM60 and pBM61, which differ only in the orientation of the multiple cloning sites within the his-3 fragment (Figure 1). These plasmids were built by first constructing plasmid pBM59 by ligating the 5.22 kb HindIII - BamHI 5’ fragment of pRAUW122, containing the 5’- truncated his- 3fragment (“left flank”; Aramayo and Metzenberg, 1996, Fungal Genet. Newsl. 43:9- 13), into pSP72 (Promega, Madison, WI) linearized with HindIII and BglII. To construct plasmids pBM60 and pBM61, the multiple cloning site from pBluescript KS(+) was amplified by PCR using Taq polymerase (Promega, Madison, WI) and ligated into SmaI- digested pBM59. The ligations took advantage of the single A overhangs left by Taq polymerase on the PCR product and single T overhangs added to pBM59 by Taq polymerase after digestion with SmaI. Both plasmids can be obtained from the Fungal Genetics Stock Center.
pBM60, pBM61 and derivatives with various test sequences inserted into the multiple cloning site were targeted to the his-3 locus by transformation of spheroplasted conidia (Aramayo and Metzenberg, 1996, Fungal Genet. Newsl. 43:9-13) or by electroporation of strains containing his-3 allele 1-234-723 (e.g., FGSC strain #6103). Electroporation of freshly harvested conidia was based on a published protocol (Vann, D.C., 1995, Fungal Genet. Newsl. 42A:53). For highest efficiency, conidia were harvested from 10 to 28 day cultures, washed and centrifuged three times in 50 ml of 1 M sorbitol, and the suspension adjusted to 2.5 x 109 conidia/ml. Linearized plasmid DNA (300 - 2000 ng) was added to 40 ul of washed conidia (1 x 108 conidia) in a 0.2 cm electroporation cuvette (BioRad Laboratories, Hercules, CA) and kept on ice for 5 min. After electroporation (voltage gradient: 1.5 kV/cm; capacitance: 25 uFD; resistance: 600 ohms; BioRad Gene Pulser, BioRad Laboratories, Hercules, CA), 1 ml of 1 M sorbitol was added to each cuvette, the suspension was mixed by gently pipetting up and down, and 250 ul was directly plated on medium containing appropriate supplements. Transformants were picked after 3 to 5 days of incubation at 32oC. Correct integration was verified by Southern analysis.
Results and Discussion. Gene targeting by transformation of spheroplasted conidia with pBM60- derived plasmids resulted in low overall transformation efficiency (approximately 2 to 15 transformants/ug of DNA) with 60% of the transformants having test sequences correctly targeted. Approximately 20% of all transformants showed wild type his-3 bands in Southern analyses and did not contain the various test sequences; approximately 20% of all transformants showed complex restriction patterns reflecting complex integration events.
Electroporation gave considerably higher numbers of His+ transformants and higher targeting efficiencies. The highest number of transformants per transformation was attained with 300 ng of plasmid DNA. Using 300 ng of DNA, we obtained from 280 to 4340 His+ transformants/ug DNA. Transformation efficiency was significantly reduced when conidia where stored in 1 M sorbitol at 4oC for 4 days (156 transformants/ug DNA) or 7 days (59 transformants/ug DNA). The frequency of gene replacements including the test sequences was high (50-100%; 71% average) in transformations with fresh conidia, but was reduced when conidia that had been stored at 4oC were used for transformation (average 40%).
In our hands, most transformants obtained were heterokaryotic, containing both transformed and untransformed nuclei, using plasmids derived from either pBM60 or pRAUW122. Therefore, when homokaryotic transformants were desired, we prepared microconidia from the primary transformants (Pandit and Maheshwari, 1994, Fungal Genet. Newsl. 40:64-65).
Acknowledgments. We thank Jane Yeadon and David Catcheside for sharing materials and unpublished results and Layne Huiet for help in translating electroporation parameters for use with the BioRad Gene Pulser. This work was supported by U.S. Public Health Service grant GM #35690 and a grant from The Human Frontiers Science Program.

Figure 1. Map and restriction sites of pBM61.
A. pBM61 is shown with all recognition sites for single- cutting restriction endonucleases indicated. Both the 5'- truncated and 3' his-3 exons and the single his-3 intron are indicated. The inner circle shows the sources of the fragments used to build pBM61, as well as vector primer sites and fusion junctions. The pBluescript KS(+) multiple cloning site is expanded below the map with recognition sites for single-cutting restriction endonucleases and primer sites indicated. Sites within the multiple cloning site that are duplicated elsewhere in the plasmid are not shown. B. All restriction endonucleases that cut pBM61 only once are listed along with their positions on the map. Coordinates run clockwise with position one at the SP6 primer site. C. All restriction endonucleases cutting only within the pSP72 fragment are indicated with their positions and number of cuts. These enzymes are useful for linearizing the plasmid in preparation for transformation.
Return to the FGN 44 Table of Contents