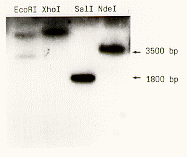
Figure 1. Southern blot analysis using the cloned TEF1 gene as a probe. DNA was digested with restriction enzymes as indicated.
Preparation methods for both DNA and RNA of filamentous fungi often are labour intensive and unreliable. The rapid preparation protocol presented here has been found to work for Schizophyllum commune and other filamentous fungi.
To prepare nucleic acids that suit molecular biologists needs, either DNA or RNA, mostly very labour intensive protocols are applied. Particularly in filamentous fungi, purification procedures may be complicated by the presence of carbohydrates excreted from the fungal cells during submerged growth. We therefore developed preparation protocols for DNA or RNA extraction from the filamentous fungus Schizophyllum commune which rely on the immediate use of fungal mycelium grown on solid media. Our protocol yields sufficient amounts of nucleic acids pure enough to be used in Southern and Northern analyses and PCR or RT- PCR, respectively.
In order to avoid excessive production of carbohydrates often observed in submerged cultures, we routinely use surface cultures of S. commune. A growing colony of approximately 3 cm2 was cut from the agar medium (complete yeast medium or minimal media) and ground in liquid nitrogen to break the cell walls. DNA extraction buffer (100 mM tris-HCl pH8.0; 10 mM EDTA; 1 % SDS) was added (500 ul to approximately 200 mg ground culture per microfuge tube), mixed briefly and incubated for 1 h at 65 C. The supernatant resulting from a 15 min centrifugation was extracted with neutral phenol/chloroform/isoamylalcohol (25:24:1), re- extracted twice with chloroform/ isoamylalcohol (24:1) and precipitated with 2.5 volumes ethanol in 0.3 M NaAcetate. The pellet was washed with 75 % ethanol and dried. If desired, RNAse was added to the resuspended DNA (100 ul in TE: tris/HCl pH8.0 10 mM; EDTA 1 mM) at 1 mg/ml and incubated at 37 C for 1 h. Extraction of RNAse was performed with phenol/chloroform/isoamylalcohol and re- extraction with chloroform/isoamylalcohol was followed by precipitation with 0.6 volumes isopropanol. The pellet was washed twice in 75 % ethanol. The DNA (20 to 250 ug) was vacuum dried and resuspended in H2O to a concentration of 1 mg/ml.
The reliability of our protocol was tested in a PCR- based approach to amplify genomic DNA that contains a fragment of the gene encoding the translation elongation factor, EF1 .
To amplify the EF1 DNA from genomic preparations, 1.5 ug of DNA were used in a PCR reaction. One single PCR fragment could be detected which was blunt end ligated into pBluescript. This cloned TEF1 fragment was used to probe genomic DNA of S. commune to show that purity of this DNA enables restriction digestion (Figure 1).

Figure 1. Southern blot analysis using the cloned TEF1 gene as a probe.
DNA was digested with restriction enzymes as indicated.
For RNA preparation a growing colony is ground in liquid nitrogen as mentioned above. To avoid the contamination with agar medium the colony can be grown on a wetable cellophane membrane (Du Pont Cellophane film 150 PD; Alba Einmachhaut, Gehring & Neiweiser GmbH, Bielefeld). The extraction then follows essentially the protocol given by Lucas et al. (1977 J. Bacteriol. 130: 1192- 1198). One ml of RNA extraction buffer (equal volumes of 150 mM Na-Acetate/4 % SDS and phenol/ chloroform/isoamylalcohol 49/49/1) is added to approximately 20 mg ground mycelium in a microfuge tube and the solution is incubated for 15 min on ice. After centrifugation for 5 min at maximal speed the upper phase is re- extracted with chloroform/isoamlyalcohol 24/1 and then immediately precipitated with 2.5 vol ethanol. RNA prepared by this method was used in northern blot analysis to detect the abundant transcript of the TEF1 gene (Figure 2).
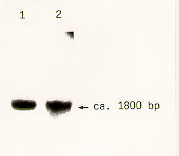
Figure 2. Northern blot analysis using TEF1 as a probe. Total RNA from
two independent preparations was separated (lanes 1 and 2, respectively).
Acknowledgements
This work was supported by the Deutsche Forschungsgemeinschaft.
Return to the FGN43 Table of Contents
Last modified 7/25/96 KMC