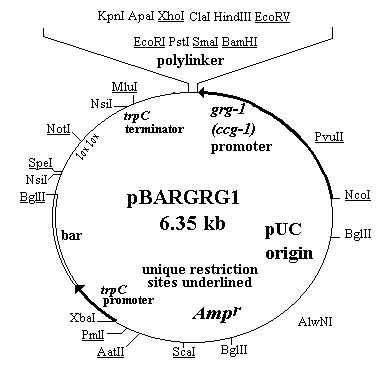
Figure 1. Fungal expression vector carrying the grg-1 (ccg-1) promoter. This vector also carries a sequence for Ignite/Basta-resistance (bar) that functions in fungi and in E. coli and the lox-NotI-lox sequence for site-specific recombination.
In the previous Fungal Genetics Newsletter, we described a series of plasmid vectors constructed carrying the bar gene as a selectable marker for use in filamentous fungi (Pall and Brunelli 1993 Fungal Genetics Newsl. 40:59-63; Pall 1993 Fungal Genetics Newsl. 40:58). In this note, we describe an additional plasmid expression vector carrying this selectable marker and the construction of four llambda/plasmid hybrid vectors carrying the bar gene within plasmid inserts that can excise by Cre/lox-mediated excision. A Neurospora crassa genomic library constructed in one of these lambda/plasmid hybrid vectors is also described below.
We have inserted the grg-1, glucose-repressible promoter (McNally and Free 1989 Curr. Genet. 14:545-551) as a 1.4 kb NcoI to BamHI fragment from pDF255, into a bar- containing expression vector in order to provide a regulatable promoter. The vector produced was designated pBARGRG1 (Fig. 1). The grg-1 gene has recently been shown to be identical to the ccg-1 clock controlled gene (S. Free, personal communication; Loros and Dunlap 1991 Mol. Cell. Biol. 11:385-388) so this promoter is also regulated by the circadian rhythm clock. It should be noted that this promoter has been patented (State University of New York) and is licenced to the Hawaii Biotechnology Group so permission must be obtained in order to use it for commercial purposes. As in the case of our other expression vectors, this one is designed to express coding sequences under their own ATGs. The pBARGRG1 plasmid is larger than those described earlier (Pall and Brunelli, op. cit.) because a larger promoter fragment is necessary in order to maintain the glucose repression (S. Free, personal communication).
This vector and the pBARGEM7-2, pBARMTE1 and pBARGPE1 vectors all contain a lox-NotI-lox fragment, as described earlier, allowing these to be attached to lambda arms, generating lambda/plasmid hybrid vectors in which the plasmid sequences can be efficiently excised by Cre/lox-mediated site-specific recombination (Brunelli and Pall 1993 Yeast 9:1309-1318; Pall and Brunelli, op. cit.). The lambda arms used for this are the same as those used previously (Brunelli and Pall, op. cit.) but a novel method of plasmid insertion was used that will be described elsewhere (Brunelli and Pall, BioTechniques, in press). All four of these lambda vectors have been constructed, and are designated as shown in Table I.
Table I. Plasmids inserted into lambda/plasmid hybrid vectors
Plasmid insert designation
pBARGEM7-2 BARGEM7
pBARMTE1 BARMTE1
pBARGPE1 BARGPE1
pBARGRG1 BARGRG1
The structure of lambda BARGEM7 is shown in figure 2.These vectors when infected into an E. coli lysogenic for lambda KC (Elledge et al. 1991 Proc. Natl. Acad. Sci. USA 88:1731-1735), carrying the cre gene and a kanamycin resistance marker, excise the plasmid inserts by Cre-lox-mediated recombination (automatic subcloning). The plasmid that is excised differs from the initial plasmid in that it only contains one lox site. As suggested earlier, we designate single lox plasmids with a prime to distinguish them from the double lox plasmids, i.e., pBARGRG1'. Another lambda/plasmid hybrid vector designed for use in Aspergillus nidulans was described recently by Holt and May (Gene 133:95-97, 1993).
It should be noted that the EcoRI and XhoI sites of the polylinker sequences of these plasmids and of the lambda vectors containing them are unique, allowing them to be used for construction of lambda genomic or cDNA libraries. A Neurospora crassa genomic library was constructed using DNA from wild type strain 74OR-23-1A as follows:
Neurospora DNA was subjected to partial digestion using the restriction enzyme Tsp509I (New England Biolabs), an enzyme that restricts 5' to the sequence AATT. This produces compatible single stranded ends to that produced by EcoRI cut DNA. Lambda DNA isolated from BARGEM7 was ligated to form concatemers, cut with EcoRI, and phosphatased using calf intestinal phosphatase. The partially restricted Neurospora DNA was run on a low melting point agarose gel by electrophoresis and the DNA was separated by size and isolated using gelase. The Neurospora DNA was then ligated to the lambda DNA and packaged using commercial packaging mix (Stratagene). A library containing approximately 2.5 X 10^6 plaques was produced with a background of 3% blue plaques (not containing inserts). When individual members of the library were excised as plasmids and restricted to determine the size of the inserts, inserts ranging from 2 kb to over 10 kb were found, averaging about 5 to 6 kb. It should be noted that this vector can package inserts of up to 11 kb in size, so these inserts are averaging about half the maximum possible size. With an estimated average insert size of 5.5 kb and Neurospora crassa genome size estimated at about 40 megabases (Perkins, 1990 Fungal Genetics Newsl. 37:9), this library should contain each Neurospora DNA sequence about 340 times.
Plasmid and lambda vectors and the Neurospora genomic library are all available from the Fungal Genetics Stock Center. We thank Dr. Steve Free for supplying plasmid pDF255 containing the grg-1 promoter. This research was supported by the College of Sciences at WSU.

Figure 1. Fungal expression vector carrying the grg-1 (ccg-1) promoter. This vector also carries a sequence for Ignite/Basta-resistance (bar) that functions in fungi and in E. coli and the lox-NotI-lox sequence for site-specific recombination.
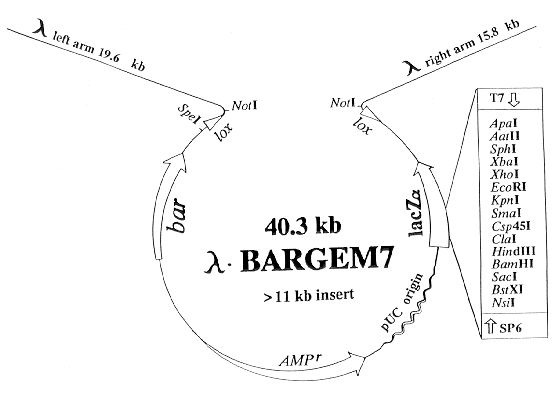
Figure 2. Diagram of the BARGEM7 vector. The restriction sites EcoRI and XhoI (in the polylinker)
and SpeI are unique in BARGEM7. Cre-mediated recombination between the two lox sites adjacent to the two arms releases the plasmid sequence. Inserts of up to 11.5 kb can be inserted into this vector and be packaged into coats. The vectors lambda-BARMTE1, lambda-BARGPE1 and lambda-BARGRG1 are similar except that they lack blue/clear screening from lacZalpha complementation and possess fungal promoters and terminators for expression of polylinker-inserted sequences in fungi.