ubi : ubiquitin
Unmapped.
Cloned and sequenced: Swissprot UBIQ_NEUCR, EMBL/GenBank U01220, X13140, NCUBQ, PIR S05323, UQNC; EST NP3F9.
Structural gene for polyubiquitin (2036).
ubi ::crp-6: ubiquitin/cytoplasmic ribosomal protein-6
Unmapped.
Cloned and sequenced: EMBL/GenBank U01221, X15338, PIR UQNCR, GenBank NCUBICRP6.
Encodes ubiquitin fused to ribosomal protein S27a. The regulatory region appears to be shared with crp-5, which encodes ribosomal protein S26. Transcription is divergent, and the two genes are expressed coordinately (2056). Called ubi-3 (2037).
uc : uracil salvage (or uracil uptake)
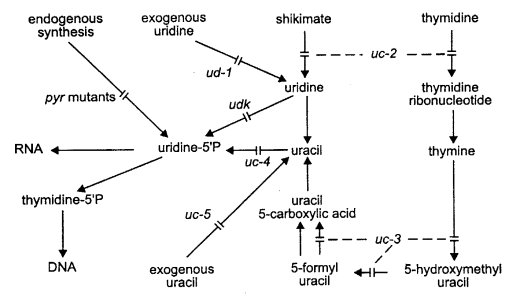
Used for mutants affecting the thymidine salvage pathway (uc-1, -2, -3, -4) or the transport of pyrimidine bases (uc-5) (2217) (Fig. 65). The symbol pyr is used for genes in the biosynthetic pathway.

FIGURE 65 The uracil salvage pathway, by which many exogenously supplied pyrimidines are cycled via uracil to uridine 5'-phosphate and thus into the latter part of the pyrimidine biosynthetic pathway. The figure shows the sites of action of the salvage-pathway uc genes: the ud-1 and uc-5 genes, which control the uptake of exogenous uridine and uracil, and udk, which controls uridine kinase (265 , 791 , 1211 , 1897 , 2217 ). For endogenous pyrimidine synthesis, see Fig. 53. From ref. (1596 ), with permission from the American Society for Microbiology.
uc-1 : uracil salvage-1
II. Linked to pyr-4 (31%) (2215, 2217).
Altered thymidine salvage pathway (Fig. 65). Able to use thymidine, thymine, 5-(hydroxymethyl)uracil, or 5-formyluracil as the sole pyrimidine source in germinating conidia of pyr-4, which is blocked in pyrimidine biosynthesis. The double mutant uc-1+ pyr-4 can use these compounds only if a primer of uridine or cytidine is supplied (2217). uc-1 causes elevated activities of enzymes that oxidatively demethylate thymine, including iso-orotate decarboxylase, and thus it is a putative regulatory gene in thymidine salvage (791, 1927).
uc-2 : uracil salvage-2
I. Linked to mat (0/12 asci) (2215, 2217).
Thymidine salvage pathway defect (Fig. 65). Unable to use thymidine or deoxyuridine as the sole pyrimidine source for pyr-4 in the presence of uc-1, but can use thymine, 5-(hydroxymethyl)uracil, 5-formyluracil, uracil, or uridine (2217). Reduced activity of the 2'-hydroxylase reactions thymidine ® thymine ribonucleoside and deoxyuridine ® uridine (1897).
uc-3 : uracil salvage-3
Unmapped.
Thymidine salvage pathway defect (Fig. 65). Unable to use thymidine, thymine, or (hydroxymethyl)cytosine to support the growth of pyr-4 in the presence of uc-1, but can use deoxyuridine, 5-formyluracil, uracil, or uridine. Excretes thymine into the medium when grown in the presence of thymidine (2217). Lacks thymine 7-hydroxylase (EC 1.14.11.6), which catalyzes the reactions thymine ® 5-(hydroxymethyl)uracil ® 5-formyluracil ® uracil-5 carboxylic acid. Apparently there is an alternate enzyme for 5-formyl uracil ® uracil-5-carboxylic acid. Apparently there is an alternate enzyme for 5-formyluracil ® uracil-5-carboxylic acid; this would explain the growth on 5-formyluracil (1211, 1897).
uc-4 : uracil salvage-4
VR. Between inl (12%) and his-6 (11%) (265). [Not in I, as was reported (2215, 2217) on the basis of 1T/8 asci with mat.]
Defective in the thymidine salvage pathway (Fig. 65). Deficient in phosphoribosyl transferase (EC 2.4.2.9) (267). Unable to use thymidine, thymine, 5-(hydroxymethyl)uracil, 5-formyluracil, or uracil to support pyr-1, even in the presence of a uridine primer (2217). The uc-4 single mutant converts 50% of supplied uridine to uracil and excretes it into the medium (2217). Resistant to 5-fluorouracil (265).
uc-5 : uracil uptake-5
IVR. Right of cot-1 (32%) (265).
Apparently defective in the transport of pyrimidine bases (Fig. 65). Unable to use any free pyrimidine base to support the growth of the pyrimidine auxotroph pyr-1, although it can use both ribose and deoxyribose nucleosides (2217). Resistant to 5-fluorouracil (265). The pyr-1 uc-5 double mutant has been used to study uptake inhibition of structural analogs (441). Selected by the inability to use uracil to supplement pyr auxotrophs or by resistance to 5-fluorouracil on ammonia-free minimal medium (265). Uracil uptake is decreased by NH3 (1245) or other good nitrogen sources or by a nit-2 mutation (266). Called tru.
ud-1 : uridine uptake-1
VIIR. Between met-7 (27%) and arg-10 (10%) (265).
Unable to use pyrimidine nucleosides to support the growth of a pyrimidine auxotroph (pyr-1) although it can use any free pyrimidine base. Probably defective in pyrimidine nucleoside transport (2217) (Fig. 65). Apparently also defective in purine nucleoside transport (440). Resistant to 5-fluorodeoxyuridine and 5-fluorouridine (265). Resistance is recessive in heterokaryons. Shows interallelic complementation (267). Scored by spotting conidial suspension on medium containing 4 ´ 10-5 M fluorodeoxyuridine, filter-sterilized (885). Mutant CIFdUrd 7, selected by resistance and called fdu-1 (885), is allelic (265). Uridine uptake is under general nitrogen metabolite regulation (266).
udk : uridine kinase
VR. Left of uc-4 (29%) (265).
Deficient in uridine kinase (EC 2.7.1.48) (267) (Fig. 65). The udk uc-4 double mutant is resistant to 5-fluorodeoxyuridine and 5-fluorouridine (265), but udk alone is not resistant to any analogue.
ufa-1 : unsaturated fatty acids-1
IVR. Linked to met-5 (7%) (1593).
Lacks stearate desaturase activity. The requirement is satisfied by 16- or 18-carbon fatty acids having a double bond at either the Δ9 or Δ11 position or 16 carbons with a trans-double bond at Δ9 or multiple cis- double bonds interrupted by methylene bridges (1849, 1850). Reverts readily, and reversions overgrow the culture on suboptimal supplement (232). Tween 80 (0.1%) is satisfactory for maintenance. Stock viability is better on Tween 80 than on fatty acids suspended in the detergent Tergitol NP-40 (1850). For the biosynthetic pathway, see ref. (1850) and Fig. 17.
ufa-2 : unsaturated fatty acids-2
IV or V. Linked to T(R2355), inl in cross to alcoy (1850).
Lacks stearate desaturase (EC 1.4.99.5) (Fig. 17). Also affects auxiliary electron transport reactions. The requirement is satisfied by 16- or 18-carbon fatty acids having a cis-double bond at either the Δ9 or Δ11 position or 16 carbons with a trans-double bond at Δ9 or multiple cis-double bonds interrupted by methylene bridges (1849, 1850). Linkage is similar to that of ufa-1. No intercross data. The designation as a second ufa locus is based solely on complementation with ufa-1 (1850). Highly revertible; stocks are lost readily on suboptimal medium (232).
ufa-3 : unsaturated fatty acids-3
I or II. Linked to T(4637) al-1 in cross to alcoy (757).
Lacks stearate desaturase activity. Requires an unsaturated fatty acid for growth. Highly revertible (753, 757).
ufa-4 : unsaturated fatty acids-4
I or II. Linked to T(4637) al-1 in cross to alcoy (757).
Affects fatty acid biosynthesis and, presumably, electron transport. Highly revertible. Growth is very slow even with fatty acid supplementation (757).
un : unknown
Used for conditional mutants for which the function is unknown. The name was introduced by early Neurospora workers for temperature-sensitive conditional mutants that are irreparable by supplementation at the restrictive temperature (34ºC or higher). Originally referred to as "unknown requirement" on the unfounded supposition that the mutants would prove to be reparable auxotrophs. Some heat-sensitive genes have been characterized sufficiently before naming to be assigned more specific names, such as ndc, rip, psi, eth-1, and fs-2. Certain heat-sensitive mutants with altered morphology at restrictive temperature have been named cot or scot. Little is known of the molecular or cellular basis of most un mutants. At least some are deficient in amino acid transport (1029, 2089). The un mutants have been useful as genetic markers. Scoring may require growth comparisons at two temperatures, preferably using small conidial inocula. Many un mutants do not achieve normal growth rates even at the permissive temperature (usually 25ºC). Temperature-sensitive auxotrophs with conventional requirements sometimes have been classed initially as un because complex complete medium is either inadequate or inhibitory. For example, among mutants called un in a study in ref. (951), nos. 3, 13, and 14 proved to be temperature-sensitive thr auxotrophs, nos. 6, 20, and 30 were his, and nos. 19 and 35 were asn (962). Locus numbers were assigned to the first eight un mutants in ref. (1586). The letter T in allele numbers of numerous un mutants designates Tokyo, not translocation.
un-1 : unknown-1
IR. Between cr-1 (5%; 9%) and nd, T(4540)R. Linked to cys-9 (0/72), bs (9%) (1578, 1582, 1592, 1603).
Function unknown. Heat-sensitive. No growth at 34ºC (909). Linear growth is suboptimal at permissive temperature (1392). Reported deficient in mitochondrial superoxide dismutase SOD2 (1392). Called un(44409).
un-2 : unknown-2
IR. Between sn, os-4, T(AR173)L and T(AR190), his-2 (1%) (1578).
Function unknown. Heat-sensitive (907), growing at 25ºC but not at 39ºC. Scorable but leaky at 34ºC. Called un(46006).
un-3 : unknown-3
Mutant 55701, first called un(55701) and then un-3, was shown to have a complex phenotype with both a defect in growth, which is temperature-sensitive, and a defect in transport, which is not. The heat-sensitive growth defect is allelic with an independently described mutation, cyt-20. The transport defect confers resistance to ethionine and p-fluorophenylalanine. The lesions responsible for temperature sensitivity and analogue resistance may be separable by recombination (1104). When ascospores from un-3+ ´ un-3 55701 were plated on minimal medium plus ethionine at 35ºC, ethionine-resistant colonies were obtained at a frequency of 0.1% (309, 1104), suggesting but not proving that recombination occurs and that separate genes may be involved. The order of the two putative mutations relative to the mat locus was not determined. Although the symbol un-3 might be considered to have priority over cyt-20 in designating the temperature-sensitive component, we propose to retain cyt-20 for the independently arising mutation that has no transport defect and to continue using un-3 for the original 55701 mutant, which has long been treated as a single gene and which will undoubtedly continue to be useful as a marker. If the ethionine-resistance trait proves to be due to mutation in a separate gene, an appropriate name for the second component would then be eth-2, and the 55701 genotype might be given explicitly as cyt-20 eth-2. The following description is based on strains of the original 55701 mutant, which is defective in both functions.
IL. Between ser-3 (1%), In(NM176)L and mat (£ 0.1%) (917, 1470, 1578). No recombination (0/21,200) or complementation with cyt-20 allele 1 (1322).
Cloned and sequenced: Swissprot SYV_NEUCR, EMBL/GenBank M64703, GenBank NEUXXX.
Includes the structural gene for valyl-tRNA synthetase mitochondrial precursor. Growth is heat-sensitive (909). Unlike cyt-20 (1631), growth of un-3 (55701) ceases sharply between 28.5 and 30ºC (1322), and there is no gross deficiency in cytochromes b and aa3 or in protein synthesis, suggesting that valyl-tRNA synthetase may have another function in addition to protein synthesis (1104). The mutant is multiply transport-deficient at permissive temperatures: resistant to ethionine and p-fluorophenylalanine at 25ºC (1028, 1029), with increased fragility of protoplasts (1029) and a reduced rate of uptake of citrulline (2089) and aspartate (1029, 2230). Used to tag the mat genes (916) and as a flanker in an attempt to resolve mat into components by recombination (1470). Used in cloning mat A (746, 2167). Mutations that are probably un-3 alleles are selected as citrulline-resistant mutants of pyr-3 arg-12s; most mutants selected in this way show complementation between alleles (2089). Growth at 25ºC is aided by 0.3 mg/ml sodium acetate. May be scored by slow growth at 25ºC on minimal medium without acetate (916). See cyt-10.
un-4 : unknown-4
VIL. Between lys-5 (2%; in a common cosmid) and T(T39M777), un-13 (3%) (1578, 1815, 1986).
Cloned by complementation (1815).
Function unknown. Heat-sensitive, 34 vs 25ºC (906). Called un(66204).
un-5 : unknown-5
IL. Between fr (6%), T(OY330)L and T(39311)L, nit-2 (2%) (1555, 1592).
Function unknown. Heat-sensitive, 34 vs 25ºC. Inhibited by histidine and tryptophan. Osmophilic at 26-28ºC (1117, 1118). Called un(b39).
un-6 : unknown-6
IIIR. Between sc (21%) and leu-1 (1546, 1582, 1592).
Function unknown. Heat-sensitive, 34 vs 25ºC (909). Called un(83106).
un-7 : unknown-7
IR. Between cyh-1, T(STL76) and al-1 (3%) (1578, 1586, 1603).
Cloned: Orbach-Sachs clones G14C06, G18H05.
Function unknown. Heat-sensitive, 34 vs 25ºC (947). Dies slowly at restrictive temperature (952). Allele T35M50 is mutant 31, called TS31 in refs. (951) and (952).
un-8 : unknown-8
IVR. Linked to pyr-1 (0/47). Between T(ALS159) and col-4 (5%); hence, right of psi-1 (1578, 1582, 1586).
Function unknown. Heat-sensitive (951). No growth at 34ºC. Morphology is abnormal at 25ºC, unlike psi, which is closely linked. Allele T27M9 is mutant 1 in ref. (951).
un-9 : unknown-9
VR. Between pyr-6 (3%) and oak (7%), his-6 (5%; 9%) (1593, 1603).
Function unknown. Growth is heat-sensitive, 34 vs 25ºC (951). Allele T54M96 is mutant 42 in ref. (951).
un-10 : unknown-10
VIIR. Between frq (9%) and arg-10, nt (22%) (1582). Right of for, frq, oli (by chromosomal walk) (1094). The order wc-1 un-10 oli frq for, shown previously, is apparently incorrect.
Cloned (1815).
Function unknown. Heat-sensitive, 34 vs 25ºC (951). Allele T42M45 is mutant 11 in ref. (951).
un-11 : unknown-11
VR. Left of his-1 (<1%) (1322).
Function unknown. Growth is heat-sensitive, 34 vs 25ºC (951). Allele T42M30 is mutant 10 in ref. (951).
un-12 : unknown-12
IVR. Linked to pdx-1 (5%), col-4 (0/73) (1603).
Function unknown. Heat-sensitive, 34 vs 25ºC (951). Allele T51M118 is mutant 17 in ref. (951).
un-13 : unknown-13
VIL. Between lys-5 (4%), un-4 (3%), ylo-1 (2%; 38 kb), T(IBj5)L and cpc-1 (1% or 2%; 105 kb), T(IBj5)R (1578, 1603, 2182).
Cloned by complementation and partially sequenced (both genomic and cDNA) (1815, 2182): Orbach-Sachs clone G55F.
The sequence suggests that un-13 encodes a tRNA synthetase (1815). Heat-sensitive, 34 vs 25ºC (951). Allele T42M24 is mutant 9 in ref. (951).
un-14 : unknown-14
IIIR. Between acr-2, thi-4 (8%; 20%) and met-8, leu-1 (5%) (1546, 1582, 1603).
Function unknown. Heat-sensitive, 34 vs 25ºC (951). Allele T54M55 is mutant 36 in ref. (951).
un-15 : unknown-15
IIR. Between trp-3 (10%) and rip-1 (1%) (1546, 1582, 1586).
Function unknown. Heat-sensitive, 34 vs 25ºC (962). Grows poorly at permissive temperature. rip-1 is therefore preferred to un-15 as a marker for the right end of II (1584). Semidominant at 39ºC. Transfers grow after 4 days at 34ºC (1546). Allele T54M50 (962).
un-16 : unknown-16
IL. Between mat (<1%) and ta (1%), acr-3 (<3%) (1582, 1603).
Function unknown. Heat-sensitive, 34 vs 25ºC (951). Allele T42M69 is mutant 16 in ref. (951).
un-17 : unknown-17
IIIR. Between tyr-1 (4%) and T(UK8-18)R. Linked to nit-7 (0/63) (1546, 1578, 1603).
Heat-sensitive, 34 vs 25ºC (951). Ascospores containing un-17 are white or slow to mature (1546). Shows rapid, exponential death at 35ºC, which is averted by cycloheximide or conditions allowing no protein synthesis. Cold-sensitive and osmotic-remediable at 11ºC (1390). Phospholipid synthesis is altered (952). When grown above 34ºC, the mutant accumulates the same phospholipid intermediates as does chol-2 (753) (Fig. 18). Reported to be deficient in extracellular superoxide dismutase SOD4 (EC 1.15.1.1) (1392). Allele T51M171 is mutant 25 or TS25 in refs. (951) and (952).
un-18 : unknown-18
IR. Between R (11%), T(MD2)L, T(NM169d)L and T(MD2)R, Tip-IR (1578, 1580).
Cloned and sequenced: GenBank AB006052.
Encodes the second largest subunit of RNA polymerase I (EC 2.7.7.6). Heat-sensitive (951). No growth at 34ºC. Growth at 25ºC is substantial but not wild-type and is better on complete than on minimal medium. The period length of circadian rhythm is increased by ~2 hr at temperatures between 22 and 32ºC (1498). Allele T54M94 is mutant 41 in ref. (951).
un-19 : unknown-19
VR. Linked to al-3 (9%), un-11 (14%) (1546).
Function unknown. Growth is heat-sensitive, 34 vs 25ºC (1546).
un-20 : unknown-20
IIR. Between aro-1 (5%; 9%), ff-1 (4%) and ace-1 (15%) (210, 1582, 1592, 2044).
Function unknown. Heat-sensitive, 39 vs 25ºC. Best scored at 39ºC on minimal medium. Leaky. Some flat, aconidiate, unpigmented growth occurs even at restrictive temperature (1582). Hyphae bleed at 39ºC. Called mo(P2402t) (1592, 2044).
un-21 : unknown-21
IIIR. Between acr-2 and met-8, un-6 (1546, 1582).
Function unknown. Heat-sensitive, 34 vs 25ºC (951). Allele T53M26 is mutant 29 in ref. (951).
un-22 : unknown-22
VII. Linked to met-7 (l%), un-10 (>20%) (1582, 1992).
Function unknown. Heat-sensitive, growing at 20-28ºC, but not at 37ºC. Called un(61C), un(62C) (1992).
un-23 : unknown-23
VIR. Between trp-2 (5%; 27%) and col-18 (3/28), T(OY320), T(UK14-1), ws-1 (1546, 1578, 1992).
Function unknown. Heat-sensitive, growing at 25ºC, but not at 28ºC. Called un(64D) (1992).
un-24 : unknown-24
IIL. Between T(AR18)L, hsp70 and het-6 (0/222; 14 kb), T(AR18)R; hence, between cys-3 and T(P1869), cot-5 (1421, 1935).
Cloned and sequenced: Genbank AF171697; Orbach-Sachs cosmids G8G1 and X14C1 contain both het-6 and un-24.
Encodes the large subunit of ribonucleotide reductase (1936). A variable domain confers vegetative incompatibility when un-24OR is transformed into spheroplasts of a heterotypic strain, un-24PA (1937). Growth is heat-sensitive, 34 vs 25ºC. The growth defect at restrictive temperature is partially overcome on media of high osmotic pressure (1936). Obtained by RIP in progeny from Dp(AR18) ´ Normal sequence (976, 1589, 1936). un-24 and het-6 are in linkage disequilibrium, with no recombination coupling phase in strains from natural populations (1351).
un-25 : unknown-25
VIL. Right of cpl-1 (4%) (1815). Linked to ylo-1 (0/86) (1582).
Cloned and sequenced (1815).
The sequence suggests that un-25 encodes ribosomal protein L13. Not complemented by DNA from either un-4+ or un-13+ (1815). Heat-sensitive, 34 vs 25ºC (951). Allele T51M154 is mutant 22 in ref. (951).
Allelic with fls.
upr-1 : ultraviolet photoreactivation-1
IL. Between mat (2%) and arg-1 (7%) (2132).
Epistasis group Upr-1 (Table 3). The original allele shows moderate sensitivity to UV (494, 2133), very low sensitivity to ionizing radiation (1846), moderate or low sensitivity to MNNG, 4-NQO, ICR-170, (953) and nitrous acid (2132), and no sensitivity to MMS (1009) or histidine (1469). (See mus and uvs entries for analyses of other genes conferring mutagen sensitivity.) A second upr-1 allele has been obtained among supersensitive double mutants by mutagenizing the highly UV-mutable, slightly UV-sensitive mus-18 (960; see mus-38 for method). Homozygous fertile. Defective photoreactivation of UV damage in vivo, but the photoreactivation enzyme is functional in vitro (2132, 2133). Survival after UV irradiation is increased by photoreactivation only at UV levels giving <10% survival, as was also found for mus-26 (957), which is in the same epistasis group and which is phenotypically similar (949, 957). Earlier results, at least in tests with UV (947), had suggested that upr-1 and uvs-2 belonged to the same epistasis group. Although such a grouping was plausible considering other similarities (it figured in the literature for some time without being rechecked; 1012, 1836), results now favor placing upr-1 with mus-26 in a separate group (949, 957, 1838). A subgroup is possible.
ure : urease
Mutants at four loci, numbered ure-1 to -4, lack all detectable urease activity (Figs. 11 and 65). Possibly all four are structural genes; urease from other organisms comprises numerous distinct subunits of a hetero-oligomeric protein (156). ure mutant D2, which is tightly linked to ure-1, fails to complement mutants at any of the four loci, suggesting a regulatory role (862). Mutants of another type possess partial activity, but are readily scorable by poor growth on urea as the N-source. These probably represent additional loci in V (A7, S3), IV (E3, E7), or elsewhere (C5, K3, R2) (862, 914). They are not given separate entries here. Mutants have been isolated by methods based on the inability to generate ammonia from urea, using pH indicators (156, 862, 1087). The following methods for scoring isolates are probably generally applicable, but all have not been tested on mutants at all loci. Method 1: Little growth on filter-sterilized urea as the sole N source (862) (not good if amino acids must be added). Method 2: 5- to 20-min scoring test touching bits of filter paper, dipped in urea-bromthymol blue buffer, to conidia or aerial growth (1086). Method 3: Color change when grown on slants of synthetic crossing medium (2208) containing phenol red and urea at 3 mg/ml as the sole N source, autoclaved, and scored after 4-5 days at 34ºC (1055, 1582).
ure-1 : urease-1
VR. Between ure-2 (3%), am (1%; 150 kb), ace-5 (<1%) and his-1 (<1%) (217, 1086, 1122).
No urease activity (EC 3.5.1.5) (862, 1087) (Figs 11 and 65). Some revertants show altered heat stability of urease, suggesting a structural role (156). Used to study the metabolic fate of arginine by measuring urea accumulation (1156). Used for arginine tracer experiments (424) and flux (657). Used to determine the relative contributions of arginine and purines in urea formation (454). For the role of urease in purine catabolism, see xdh-1. For scoring methods, see ure. Possible allele D2 fails to complement ure-1, -2, -3, -4 (862). Called ure(9) (1086).
ure-2 : urease-2
VR. Between sp (8%) and am (2%), ure-1 (3%) (220, 1086).
No urease activity (EC 3.5.1.5) (862, 1087) (Figs. 11 and 65). The enzyme from interallelic complementation and from some revertants shows altered heat stability, suggesting a structural role (156). Shows hyperinducibility of purine catabolic enzymes uricase, allantoicase, and allantoinase (1712). Reversion and complementation data (156). For scoring methods, see ure. Called ure(47) (1086).
ure-3 : urease-3
IIR. Between arg-12 (7%; 12%) and un-20 (8%), ace-1 (15%) (1582). Because allele B1 is closely linked to translocation T(IR;II)B1 in the original strain, ure-3 was at first assigned incorrectly to IR (156, 862). Point-mutant allele F29 showed linkage in IIR, not IR (1582).
No urease activity (EC 3.5.1.5) (Fig. 8). Some revertants show altered urease thermostability, suggesting a structural role (156). For scoring methods, see ure.
ure-4 : urease-4
IR. Left of ad-3B (3%). Probably right of his-3 (1%) (156).
No urease activity (EC 3.5.1.5) (862) (Fig. 8). Some revertants show altered heat stability of urease, suggesting a structural role (156). For scoring methods, see ure.
ure(D2) : urease (D2)
VR. At or near ure-1 (0/45) (862).
No urease activity. Fails to complement representatives of all four ure loci; possible regulatory gene (862).
usg-1 : upstream gene-1
VR. Located 3.5-kb upstream of the distal regulatory sequence of am (670).
A gene of unknown function, transcribed in the same direction as am (670).
uve-1 : UV-endonuclease-1
Changed to phr.
uvs : ultraviolet sensitive
The uvs (and upr) mutants of Neurospora are defective in DNA repair and have highly pleiotropic phenotypes (946, 1838) (Table 3). Genes designated uvs-1 through uvs-6 were identified prior to 1980; after that time, mus (mutagen sensitive) was adopted as the symbol to designate mutants defective in DNA repair, beginning with mus-7 (1839). Thus, whereas a given mus mutant may not be sensitive to UV, any UV-sensitive mutant isolated since 1980 has been given a mus designation. The sensitivity of uvs mutants is typically recessive. Most of these mutants are also sensitive to various genotoxic chemicals that can be incorporated into agar media for tests during cell division. Scoring of qualitative differences in UV sensitivity is accomplished most readily by spot tests in which the growth of cells on exposed plates and control plates is compared (1830, 1993). For the more accurate assessment, survival curves are used, especially when single and double mutants are compared (1836). Sensitivities to different genotoxic agents may be associated in patterns and can be diagnostic for certain DNA repair types or epistatic groups (see the introduction to mus genes). In general, the increases in UV sensitivity observed for UV sensitive mutants in filamentous fungi such as Neurospora have been relatively low. This has made genetic analysis labor-intensive. Thus, uvs-2, which has the highest UV sensitivity for any single Neurospora crassa mutant, is only 20-fold more sensitive to UV than is the wild type (1833, 1838). In contrast, 100- to 200-fold increases in sensitivity are reported for mutants of Escherichia coli and Saccharomyces cerevisiae. Some of this difference is now attributed to the alternative repair pathways (e.g., mus-18 and mus-38) available in filamentous fungi (also in fission yeast and probably in higher eukaryotes). Properties of the uvs and related mutants are summarized in refs. (946), (948), (953), (1010), (1015), (1835), (1836), (1838), and (1839). For properties of double mutants, see refs. (855), (947), (957)-(960), (1012), and (1017).
uvs-1 : ultraviolet sensitive-1
Unmapped (355).
Repeated efforts have failed to confirm that uvs-1 is a single Mendelian mutant. The presumed mutant strain is therefore no longer used in investigations (1836, 1838).
uvs-2 : ultraviolet sensitive-2
IVR. Between cys-4 (5%) (1993) and pmb (8%) [S. Ogilvie-Villa, cited in refs. (513) and (1776)], T(S4342)R, T(AR209), T(T54M50), T(ALS179) (2-6%) (1578).
Cloned and sequenced: Swissprot UVS2_NEUCR, EMBL/GenBank D11458, PIR S34825, GenBank NEUUVS2.
Epistasis group Uvs-2 (Table 3). uvs-2 has the highest UV sensitivity of all Neurospora uvs mutants [see uvs and refs. (494) and (1993)]. The allele usually used was discovered in stocks of mixed ancestry and may be present in other laboratory stocks (1984, 1993). Recessive in heterokaryons (1993). uvs-2 is sensitive to ionizing radiation (1833, 1846), MNNG (953, 1833), 4NQO, nitrous acid (1984), ICR-170 (953), and mitomycin C (1012). Highly sensitive to MMS (1009), as was found for several other uvs mutants. A second allele (SA4) with similarly high UV and MMS sensitivity was isolated among mutants identified by MMS sensitivity (948). Shows slight or no sensitivity to histidine (1473). Normal spontaneous mutation and high UV-induced mutation (88, 494, 497, 502). Homozygous fertile. Meiosis and meiotic crossing over are unaffected. Used to show that the DNA repair system is induced by a small dose of UV (1983) and to demonstrate post replication repair (273).
The uvs-2+ gene was isolated by transformation of uvs-2 with a genomic library and selection for resistance on MMS medium. Encodes a structural gene homologous to Saccharomyces RAD18 (2099) and Aspergillus uvsH (2281). However, the amino acid identity with RAD18 is only 25%, and there is no functional complementation of the yeast RAD18 mutant with uvs-2 cDNA cloned in a yeast expression vector (1952). No major changes in the two-dimensional profile of polypeptides were detected when the mutant was compared to wild type under uninduced conditions; but differences in some inducible peptides were evident (911-913). Originally, the finding that most of the phenotypic properties of uvs-2 resembled those of yeast nucleotide excision repair mutants suggested that uvs-2 was involved in the excision of UV dimers. Supporting this hypothesis, no excised dimers were detectable by chromatographic tests of uvs-2 mutants (2247). Two observations were contradictory, however: the observed X-ray sensitivity of uvs-2 and the finding in uvs-2 strains (and in all tested UV-sensitive Neurospora strains) that UV-dimer repair occurs at the wild-type level when unrepaired dimers are identified by UV-dimer specific endonuclease or antibody (89, 90, 1239). These observations are consistent with the discovery that the excision of dimers in Neurospora occurs not only by the same mechanism identified for nucleotide excision repair genes in yeast and human cells but also by a UV-dimer-specific endonucleolytic mechanism that is roughly similar but distinct from that found in Micrococcus luteus (see mus-18). The homology of uvs-2 to Saccharomyces RAD18, which is active in postreplication repair (2099), and the discovery in Neurospora of two bona fide nucleotide excision repair genes, mus-38 and mus-40, which are homologous to yeast RAD1 and RAD2, respectively (855), finally provided the expected parallels with the yeast repair system. Thus, whereas uvs-2 is known not to be involved in a known process of excision, the actual function carried out by genes of the Uvs-2 epistasis group (Table 3), or by their yeast homologs RAD18 and RAD6, remains unknown.
uvs-3 : ultraviolet sensitive-3
IVL. Linked to cys-10 (3%; 7%), probably to the left (1014, 1830).
Cloned and partially sequenced (961).
Epistasis group Uvs-3 (Table 3) (1012). Sensitive to UV (494, 1010, 1830, 1831) and highly sensitive to MMS (1009, 1012). Moderately sensitive to 5-azacytidine (662). The commonly used allele ALS11 has low and variable conidial viability (502, 1830). A second uvs-3 allele, FK016 (1012), has approximately 2-fold better conidial viability and forms more uniform colonies. Both alleles are sensitive to histidine, MNNG, X rays, g rays, and mitomycin C (381, 953, 1010, 1831, 1846). However, ALS11 is significantly more sensitive than FK016 to X rays, g rays, and mitomycin C (1012). Other agents have also been tested for their effects on survival and mutation (953). uvs-3 displays a mutator phenotype (502, 1010), resulting in spontaneous reversions and variable sensitivities in specific isolates (1012, 1830). There is no induction of mutation by UV (494) or by g rays and chemical mutagens (953, 1846). Therefore, the mutant is probably defective in error-prone repair (1834). Homozygous barren; sexual development is blocked before karyogamy (1685, 1830). Morphological mutants are frequent in backcrosses and ascospore viability is low; few asci contain eight black spores. Duplication instability is increased significantly, and this is correlated with increased sensitivity to hydroxyurea (1835). Sometimes gives stop-start growth on minimal medium or minimal medium + histidine (1467). Photoreactivation is impaired in vivo, but the photoreactivation enzyme functions in vitro (960, 1832). Levels of endo-exonuclease are low, probably accounting for the reduced nuclease halo phenotype (1014). The mutant appears deficient in proteolytic conversion of nuclease precursors to active intra- and extracellular DNases, but this effect could be indirect (666). Protease activity may be reduced (1723). The mutant shows a high constitutive level of repair in rescuing a heterokaryotic component that carries potentially lethal mutagen-induced damage (1983). Excised dimers were reported to be detectable only after a delay and to be released at a reduced rate (2247). However, alternate tests indicated that, under specific conditions, constitutive levels of dimer disappearance from DNA were as high in uvs-3 as the induced level in wild type (88, 1838). One significant charge change in a polypeptide is observed in constitutive vs inducible expression of polypeptides detected in two-dimensional profiles of near-isogenic strains of both uvs-3 alleles compared to wild type (911-913). The double mutant uvs-3; uvs-6 is inviable, as are the double mutants mus-9; uvs-6 (947, 1012) and uvs-3; mei-3 (958). However, when double mutants are tested for MMS survival, a highly synergistic interaction is found for uvs-3 with uvs-2 and with mus-8 (1012). Similarly, for the uvs-3; upr-1 double mutant, reduction of UV survival is at least additive; measurements of mutation rates also indicate interaction between the two. Spontaneous mutation levels are low for uvs-3; upr-1 (resembling upr-1); there is no increase after UV induction (resembling uvs-3) (947). These results establish the Uvs-3 group (including mus-9) separate from the Uvs-2 group (including mus-8) and separate from the Uvs-6 group (including mei-3). Additive (or stronger) interactions with uvs-3 have also been demonstrated for mus-18 and mus-38, which interact synergistically with each other and which represent different epistatis groups and repair types (959, 960). Allele FK016 was isolated as a nuclease halo mutant and originally called nuh-4 (1014). A strain used in ref. (667) and incorrectly called nuh-4 was actually uvs(FK104), a mus-9 allele. This error explains what appeared to be contradictions concerning uvs-3 phenotypes (1012).
uvs-4 : ultraviolet sensitive-4
IIIR. Left of ad-4 (4%) (1830).
Epistasis group Uvs-2 (1838) (Table 3). Sensitive to UV (494, 1830, 1831) and histidine (1467, 1835); moderately sensitive to MMS; slightly sensitive to MNNG (1010, 1831) and no increased sensitivity to ionizing radiation (1846) or mitomycin C (381). Gives stop-start growth (1467). Spontaneous mutation is probably normal (502); UV-induced mutation is reduced (494). For induction of mutation by other agents, see ref. (1846). Homozygous fertile, although ascospore viability and early growth are severely impaired in homozygous crosses (1830). No effect on mitotic gene conversion or recombination (1829). Chromosome stability in duplication strains is normal (1835). Defects in mitochondrial DNA (856). Difficult to work with. Survival of conidia is poor, even on silica gel. Recessive in heterokaryons. Stocks are best maintained in a heterokaryon (1576).
uvs-5 : ultraviolet sensitive-5
IIIR. Linked to vel (1%) (1830).
Sensitive to UV (494, 1830, 1831), MNNG, and ICR-170 (953). Not sensitive to ionizing radiation (1846), 4NQO (953), or histidine (1467). Spontaneous mutation is normal (502); UV-induced mutation is reduced (494). For induction of mutation by other agents, see refs. (953) and (1846). Recovery from heterozygous crosses is low. Meiotic recombination is not affected. Homozygous barren (1830), with meiosis blocked at pachytene (1685). No increase in mitotic chromosomal instability (1835). Abnormal mitochondrial DNA (856). Difficult to work with. Growth is slow (1838). Consequently, uvs-5 is no longer used in investigations. Stocks are best maintained in a heterokaryon (1576).
uvs-6 : ultraviolet sensitive-6
IR. Linked to met-6 (<1%). Between thi-1 (3%; 8%) and ad-9 (4%) (1476, 1837).
Epistasis group Uvs-6 (828, 958, 1012) (Table 3). Reduced conidial viability (1010). Switches to stop-start growth after initial normal growth (1467). Not completely recessive in heterokaryotic conidia (1900). Increased sensitivity to UV (494, 1837, 1846), MNNG (953, 1010), and histidine (1466, 1467). Also sensitive to ICR-170, 4NQO (953), MMS, and possibly mitomycin C (1010). Spontaneous mutation is normal in the ad-3 haploid mutation system (502); a 2-fold increase in spontaneous mutation found for recessive lethals in a heterokaryon test system was not statistically significant and may have resulted from an unusual chance occurrence of several large clones (1013). UV-induced mutation is normal in both mutation test systems (494, 1013). For mutation induction by other agents, see refs. (953) and (1846). Homozygous barren, with a block at crozier differentiation (1685). Increased duplication instability (1473). The induction of a specific polypeptide in response to UV is altered compared to wild type, as was also found for mei-2 and mei-3 (911-913). Possibly defective in nuclease levels or in secretion, giving reduced nuclease halos on DNase test medium (667, 1014); DNase A is much reduced in culture filtrates of cells grown in sorbose + DNA medium (see nuh for nuclease assays). Protease activity may be reduced (1723). Double mutants of uvs-6 with the two uvs-3 alleles ALS11 and FK016 are inviable (947, 1012). Double mutants with representative members of other epistasis groups showed additive or synergistic interactions, however (947, 957, 959, 960). Homology of at least two genes of the Neurospora Uvs-6 epistasis group with genes of the yeast Rad52 group suggests that the function may be in recombinational repair (828, 853).
uvs(FK104) : ultraviolet sensitive (FK104)
A mus-9 allele (1014). Mislabeled as nuh-4 in ref. (667).
Return to the 2000 Neurospora compendium main page