Use of a bacterial Hygromycin B resistance gene as a dominant selectable
marker in Neurospora crassa transformation.
Fungal Genetics Newsl. 36:79-81 (1989)
Staben, C.(1), B. Jensen(2), M. Singer(2), J. Pollock(3), M.
Schechtman(3,4),J. Kinsey(5) and E. Selker(2) - (1)Department of
Biological Sciences, Stanford University, Stanford, CA 94305
(2)Institute of Molecular Biology, University of Oregon, Eugene. OR
97403 (3)Department of Biology, Syracuse University, 130 College
Place, Syracuse, NY 13244 (4)Present Address: USDA, APHIS,
Federal Center Building, 6505 Bellcrest Road, Hyattsville, MD 20782
(5)Department of Microbiology, University of Kansas Medical Center,
Kansas City, KS 66103.
Dominant transformation markers allow maximum flexibility in the
choice of transformation recipients. The most widely used
transformation marker in N. crassa has been the Neurospora gene
conferring benomyl resistance (Orbach, et al. 1986 Mol. Cell. Biol.
6:2456-2461). Unfortunately, this marker is usually inactivated
during sexual crosses, presumably because its introduction by
transformation generates strains with duplicate beta-tubulin
sequences that are subject to inactivation during the sexual cycle by
RIP (Selker et al. 1987 Cell 51:741-752; Selker and Garrett, 1988
Proc. Natl. Acad. Sci. USA 85:6870-6874). We therefore looked for a
dominant drug resistance marker that would not be homologous to any
sequence in the Neurospora genome. A number of researchers have
found that a bacterial gene encoding hygromycin B resistance (hph,
hygromycin B phosphotransferase), if driven by a eukaryotic promoter,
can confer drug resistance on eukaryotic cells. Hygromycin B (hygB) is
an amino-glycoside that inhibits protein synthesis by causing
mistranslation (Gonzalez et al. 1978 Biochim. Biophys. Acta 521:459-
469) and by interfering with protein translocation (Singh et al. 1979
Nature 277:146-148). In fungi, Gritz and Davies have had promising
results in the use of the hph gene in yeast (Gritz and Davies 1983
Gene 29:179-188). Punt et al. and Cullen et al. have had success using
this gene in both A. nidulans and A. niger (Punt et al. 1987 Gene
56:117-124; Cullen et al. 1987 Gene 57:21-26). Here we report the
sensitivity of Neurospora to hygB and the use of two existing plasmids
and derivatives as transformation markers in Neurospora.
All of the strains of Neurospora that we have tested (approx.
15) are sensitive to hygB. We generally use Vogel's minimal medium
with sorbose for plates or sucrose for slants. We have selected
resistance in a number of supplemented media, including a complete
medium. HygB was obtained from Calbiochem. The commercial
powder is dissolved in sterile water at 200 mg/ml and added to
autoclaved media. Media can be stored for several months at 4oC. A
number of different sensitivity tests have been performed. In general,
strains are sensitive to 150 micrograms/ml hygB. Several different
plating conditions have been used to isolate transformants produced by
standard procedures (Vollmer and Yanofsky 1986 Proc. Natl. Acad.
Sci. USA 83:4869-4873). Our standard conditions involve plating 5 x
106 spheroplasts on a 25 ml sorbose plate (200 micrograms/ml hygB)
in 8 ml top agar lacking hygB. Top and bottom layers usually contain
1.5% agar. Decreasing the volume of top agar or increasing the hygB
concentration increases the stringency of the selection and improves
the background, but discriminates against some transformants and
lowers the apparent transformation frequency. We have used similar
selection conditions to isolate hyg resistant mutants from mutagenized
conidia. The frequency of spontaneous mutation to hyg resistance is
not high enough to interfere with the selection.
Our initial transformations used either pAN7-1 (Punt et al. 1987
Gene 56:117-124) or pDH25 (Cullen et al. 1987 Gene 57:21-26).
These plasmids contain Aspergillus transcription signals that direct the
expression of the E. coli hph gene. We have made derivatives of these
plasmids that other researchers may find useful. In pES200 the trpC
terminator has been deleted from pDH25 leaving a single BamHI site
(Figure 1A). This plasmid is as effective as pDH25 in conferring hygB
resistance; the single BamHI site allows convenient cloning of BamHI
compatible restriction fragments. Two other derivatives, pCSN43 and
pCSN44, contain a SalI fragment of pDH25 that confers resistance as
well as pDH25. These plasmids provide additional unique cloning sites
(Figure 1b, 1c). They offer the additional advantage that they
replicate to a very high copy number in E. coli. The SalI fragment
from pDH25 that is present in pCSN43 and pCSN44 has been trans-
ferred into several other plasmids, and has conferred hygB resistance
in each case. We have not mapped the actual transcription signals or
start sites that direct hph gene expression in N. crassa, though the
structure-activity relationship for the trpC promoter has been studied
in Aspergillus (Hamer and Timberlake 1987 Mol. Cell. Biol. 7:2352-
2359). Surprisingly, insertion of a 430 bp TaqI fragment from the zeta
region (Selker and Stevens, 1985 Proc. Natl. Acad. Sci. USA 82:8114-
8118) into the ClaI site of pDH25, separating the promoter from the
hph coding region, did not abolish hygBr. However, the ClaI-BamHI
fragment containing the hph coding region is not sufficient for
resistance. Plasmids pES200, pCSN43 and pCSN44 are available from
the Fungal Genetics Stock Center.
Transformation frequencies with the hph gene are similar to
those observed with the benomyl resistance gene. Mixtures of
plasmids containing the two markers cotransform at frequencies up to
75%, suggesting that there is little difference between their transfor-
mation frequencies. We observe 1-10 x 103 transformants/microgram
DNA under the conditions that we have described. Transformants
appear to fall into two classes: those resistant to 200 microgram/ml
and those resistant to >1 mg/ml. We have not determined the
difference between classes. Southern hybridization of various
transformants indicates that one to several copies of the gene have
integrated, as observed for most other transforming DNAs.
Transformants typically yield hygromycin resistant progeny in sexual
crosses with wild type. The hph marker is inactivated when known
multicopy transformants are crossed to wild type
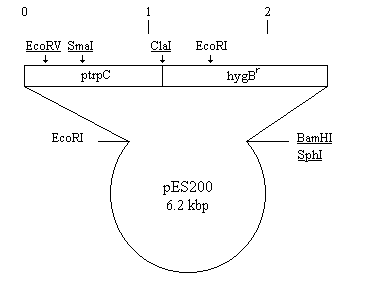
Figure 1A: pES200 was constructed by deleting the 710 bp BamHI
fragment from pDH25. Sites discussed in the text are indicated. The
position of the trpC promoter and hygBr are indicated. Unique
cloning sites are underlined. Vector sequences are represented by the
thin circle; the resistance gene and its transcription signals are drawn
to scale as a thick box.

Figure 1B: pCSN43 contains the 2.4 kbp SalI site from pDH25 cloned
into the SalI site of pBSSK+ (Stratagene). The indicated scale is in
kilobase pairs. Unique restriction sites are underlined. The promoter
and terminator fragments from Aspergillus trpC are also indicated.
Additional restriction sites in these plasmids may be unique. Those
indicated have been tested.

Figure 1C: pCSN44 has the SalI fragment from pDH25 oriented
opposite to the orientation in pCSN43.


