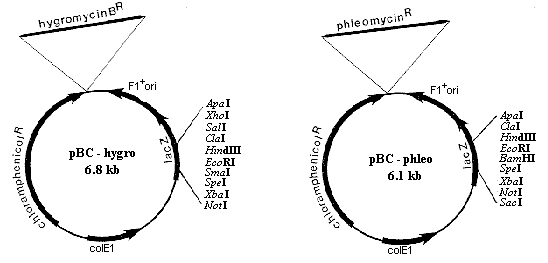
Figure 1. Two vectors for transformation of filamentous fungi. Left: pBC-Hygro. Right: pBC-phleo. For details, see text.
Several vectors containing dominant antibiotic resistance markers for the transformation of filamentous fungi are available. However, most of them lack a recombinant selection system and a polylinker containing a sufficient number of cloning sites, thereby limiting the subcloning strategies. I have constructed two new vectors, derived from the pBC-SK+ bluescript vector (Stratagene), that possess these characteristics as well as many other interesting features. The restriction maps and the available cloning sites of both plasmids are presented in Figure 1. Transformants in bacteria are selected on chloramphenicol-containing medium. Recombinants can be screened by using the blue/white selection associated with the lacZ system. The polylinkers of both plasmids contain numerous unique cloning sites and are flanked by two BssHII sites as well as T3 and T7 promoters for RNA production. The plasmids display unaltered f1 origins for the recovery of single-stranded DNA for sequencing or mutagenesis.
The first plasmid, pBC-Hygro (Figure 1) carries the hygromycin B resistance cassette included in the AvaI-SphI restriction fragment from plasmid pMOcosX (Orbach 1994 Gene 150:159-162) in which the hygR coding sequence is under the control of the Neurospora crassa cpc-1 promoter and the Aspergillus nidulans trpC terminator.
The second plasmid, pBC-phleo (Figure 1), carries the phleomycin resistance cassette included in the EcoRI-HindIII restriction fragment from pUT703 (Camels et al. 1991 Curr. Genet. 20:309-314) in which the ble gene is under the control of the Aspergillus nidulans gpd promoter and the Saccharomyces cerevisiae CYC1 terminator.
We use both plasmids to transform Podospora anserina protoplasts with standard protocols and recover in each case about 100 transformants per microgram of DNA. These plasmids are available upon request.

Figure 1. Two vectors for transformation of filamentous fungi. Left: pBC-Hygro. Right: pBC-phleo. For details, see text.